Understanding the steps in metabolic pathways is important for understanding health and disease, whether in people or plants. But figuring out these biochemical pathways is difficult and time consuming. So why not have a machine do it?
Scientists at UC Davis and Ben-Gurion University in Israel have applied machine learning (artificial intelligence) techniques to this problem in tomatoes. They were able to successfully predict new, previously unknown metabolic pathways. The work is published June 18 in the journal Communications Biology.
David Toubiana, a research specialist working with Professor Eduardo Blumwald at the UC Davis Department of Plant Sciences, began by collecting existing, known metabolic pathways from public databases. He also constructed “correlation-based networks” of metabolites – biochemicals known to be present in a line of specially-bred tomato plants. These networks link similar molecules, without saying anything about actual pathways between them.

Tomatoes from the UC Davis Student Farm. Researchers are using machine learning and artificial intelligence to discover new metabolic pathways in tomatoes.
Toubiana and colleagues mapped the known pathways on to the correlation networks to create a set of partial metabolic networks. They used these networks to train machine-learning algorithms to identify new pathways. In machine learning, a computer system “learns” a new skill without being explicitly programmed for it.
Four new pathways identified
Based on this training and raw data from analyzing all the biochemicals in tomato fruit, the machine was able to identify four entirely new pathways in tomato. They were: beta-alanine degradation-I, tryptophan-degradation-VII-via-indole-3-pyruvate (previously unknown in plants), beta-alanine biosynthesis-III, and melibiose degradation. The existence of the melibiose degradation pathway was confirmed in wet lab experiments.
The combination of correlation network analysis and machine learning is a powerful new tool for discovering metabolic pathways and networks, even when there is incomplete knowledge of all the genes and molecules involved, the researchers predict.
Additional authors on the study are: Rami Puzis, Lingling Wen, Noga Sikron, Assylay Kurmanbayeva, Aigerim Soltabayeva, Aaron Fait, Moshe Sagi and Yuval Elovici, Ben-Gurion University of the Negev, Israel; and Nir Sade and Maria del Mar Rubio Wilhelmi, UC Davis Department of Plant Sciences. The tomato plants used were bred by Dani Zamir at the Hebrew University of Jerusalem.
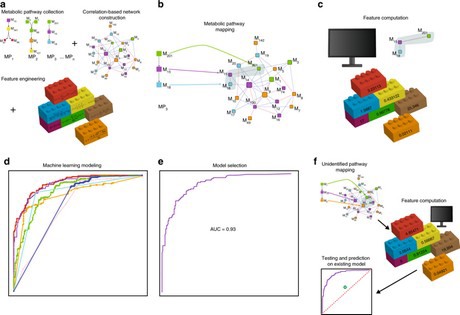
Graphic from publication in Communications Biology.
Source: UC Davis (Ann Filmer)
